Team:WalthamHS BioHawks
From 2014hs.igem.org
| Home | History of Waltham -Outreach |
Team | Project -Design |
Sponsors | Our Poster |
| Official Team Profile |
|---|
Contents |
History of Waltham
Waltham, Massachusetts was first settled in 1634. As time passed, Waltham became the forefront of the American Industrial Revolution. In the early 19th century, Francis Cabot Lowell, known as the Father of the Industrial Revolution, founded the Boston Manufacturing Company in Waltham. At his factory, located along the Charles River, he created the Waltham System, a manufacturing model that transforms raw material into a completed product at one location. Before this system, the processes of carding, spinning, weaving and fulling cotton were carried on in separate establishments under different proprietors. It was in1813 that Francis Cabot Lowell combined the processes of turning cotton into cloth under on roof, creating the world renowned Waltham System. Because of this system, Waltham became the epicenter of the American Industrial Revolution, and one of the most productive cities in America. Companies such as the Waltham Watch Company, located in Waltham, created millions of watches from the late 19th century through the first half of the 20th century. This is how Waltham got its nickname, “The Watch City”. The Waltham Watch factory was known for its engineering advancements in the standardization of parts, automation of assembly, and use of precision tools. These are all core ideas behind the concept of synthetic biology. Today, Waltham is still an appealing site for science and engineering advances, home to such companies as Raytheon, AstraZenaca and Nova Biomedical. The city has also become a center for research, innovation, and higher education, with Brandeis University and Bentley University leading the way.
Outreach
In the fall of 2013, a group of Waltham High School students applied to the Waltham Education and Beyond Foundation for a grant to support the creation of its first iGEM team. The team was awarded a grant from the Sally Elizabeth Peters memorial grant program, the first grant ever awarded directly to a student group! Sally Elizabeth Peters was a Waltham student who died at a young age. She was a "renaissance kind of student", interested in a wide range of subjects, from music to science. Her parents created this scholarship program to fund ideas that bring together diverse groups within the Waltham community that support both the arts and the sciences.
Our iGEM team is now part of the larger Waltham High community. The whole school supports our team, helping and encouraging us to strive and achieve our goals. When we interviewed students and teachers at the high school about our project, this is what we heard:
Q: How do you feel this iGEM team, dedicated to the new frontier of biology, can impact the community within Waltham High School?
-Mr. Diluzio (History teacher) : "Amazing - competition drives ingenuity - it is innovative. It's important for WHS to be on cutting edge of science."
-Mr. Leach (Math teacher) : "Awesome. This is a great thing for the school to take part in. It is an experience that's not repetitive in the classroom. An nquisitive student culture will help benefit the school as a whole. It is so nice to hear about students creating something original."
-Student Colin Holmes: "You should always try to find new things to do, it expands new opportunities. Follow your dreams."
-Student Courtney Paschal: "This program gets students to think more and reach their full potential. It can create better communication and leadership among students."
-Student Charlene Preys: "This program can help students become more involved and interested - really useful!"
Team
Our team consists of students currently enrolled in AP Biology and a few aspiring Biology students. Once we were introduced to the idea of iGEM, people who enjoy science and wanted a hands on experience in laboratories came together to form the BioHawks team. Our team consists of a 8 students, our biology teacher, and a parent who is serving as our advisor.
Rutvi Bhatt: Hi, I am Rutvi and I am a BioHawk. I love Biology and learning about stuff we experience in everyday life. I find microbiology very interesting and lab work helps me understand the concepts. I love community service and I am involved numerous awareness activities in India and Waltham. I am in charge of making the website for iGEM and helping plan the project. I look forward to presenting our project at MIT and making Waltham Proud!!
Role: In charge of designing, editing and making the website and planning and organizing work.
Micheal Perlow: I am a recent graduate of Waltham High School, in the class of 2014. Being naturally curious, especially in the sciences, I take a particular liking to biology. Last summer, I worked with a graduate student in the Biochemistry department at Brandeis University. Dealing with molecule-interactions (between RPS30B RNA and the u1 snRNP protein), we transformed bacteria and yeast to produce the desired nucleotides or polypeptides. Using that experience combined with the AP Biology class under Mrs. Maddox taken in the 2013-2014 school year, I have developed skills and experience highly adequate for use in synthetic biology. Role: Lab technician and wiki editor.
Edward Lo: Hi, my name is Edward Lo and I am currently a junior at Waltham High school. My hobbies include playing the piano and playing tennis. I decided to join to iGEM Club because it interested me in how high school kids could customize their bacteria to perform different tasks. My job on the team is to do lab work and to provide help in the wiki and other things that need to be done. Role: Lab technician and wiki editor.
Mina Antic: Hi! I’m a junior at Waltham High School, and always been interested in the sciences (the reason why I’m in the Synthetic Bio Club and why I’m working on this team). Biology is probably (read: definitely) going to be my major when I leave high school and I thought this project would be a good experience. But besides that, I enjoy playing volleyball, painting, and sleeping. Particularly sleeping, which I don’t do as much as I want to. Role: Lab technician and wiki editor.
Molly Wack: My name is Molly Wack, and I’m a junior at Waltham High School. I joined this club because I think synthetic biology is a rapidly growing field and it’s an exciting time to join it. In my free time I like to nap and eat, and I like to play basketball and lacrosse too. Role: Wiki editor.
Leon Mamish: I am Leon Mamish take out the student and what do you get? Genius, Billionaire, Playboy (ok not really), Philanthropist. But seriously, I am a student at Waltham High School interested in the sciences, specifically Biology and Chemistry. I joined the iGem Team due to my strong interest in Biology and plan to peruse a career in science. My hobbies include sleeping, watching movies, sports and eating. Also, sleeping. Role: Lab technician and wiki editor.
Jason Gonsalves: I’ve always had a peculiar interest for science. I’ve completed Environmental Science, Biology, Chemistry and AP Biology in high school and my interest has only grown stronger. When I heard about Synthetic Biology Club, I just had to check it out to sate my pursuit of the field. I love biology specifically and wish to pursue a carrier in it. My other hobbies are track and field, writing and sleeping. Role: Lab technician
Marisa Maddox: I have been teaching high school biology at Waltham for seven years. I received my Bachelor's and Master's degrees in microbiology from UMASS Amherst. In my free time I enjoy spending time with my family, traveling and sleeping. Role: Instructor
Edward Wack : I am an adviser to the Waltham BioHawks iGEM team. When I'm not coaching youth sports or gardening and doing yard work, I lead the Bioengineering Systems and Technology Group at MIT Lincoln Laboratory. Role: Advisor
Project
Throughout the fall, and into the winter, our team created a list of many possible topics for our project. We had narrowed our list down to five possible subjects: a pH sensor for tap water at our school, an environmentally friendly paper towel degrader, a stain remover for protein based stains, a pre-food taster that could detect any possible pathogens, and a more effective anti-allergy molecule. After a long discussion of the pros and cons of each topic, the team narrowed the debate to two topics: the pH sensor and the stain remover. There was much lobbying and back and forth before the majority of people voted for the stain removal project. And so began our journey.
Our project was to create an environmentally friendly alternative to the harsh chemicals in modern detergents. More and more people every day are developing negative side effects from the continuous use of the damaging chemicals. Laundry detergents can cause such side effects as rash, red and blistered skin, sun sensitivity, sneezing, and itchy or watery eyes. Our goal was to create a powerful stain remover that was safe to use for all people.
Each team member was given a part of the problem to research. Some looked at ingredients in common detergents, others researched natural stain removing remedies. In the end, we discovered a protease enzyme called subtilisin, a broad range protein degrader from the genus of bacteria Bacillus. Subtilisin is an enzyme found in many common laundry detergents and dish washers, but also found in nature. It can be obtained in certain soil bacteria, which create it in large amounts. Subtilisin is made up of approximately 269-275 amino acids, with a typical globular shape. The active site of subtilisin involves Asp-32, His-64, and most importantly Ser-221. These three amino acids converge in a 3D shape to create the active site. The Ser-221 cleaves peptide bonds with its partially negative oxygen.
Our task was to transform the subtilisin enzyme into E. coli to create a bacteria that could rapidly produce many of the needed enzymes. E. coli has a reproduction rate of 15-20 minutes, which makes it able to quickly create daughter cells. The quick reproduction rate allows the subtilisin enzyme to build up and easily break down stains.
Subtilisin also has the capability to break down blood clots. Blood clots occurs when blood platelets mature and combine with fibrin to form a liquid-like gel. The blood clots naturally in the circulatory system to prevent blood loss in damaged blood vessels. However, the coagulation of blood can be irregular, which can be serious if developed near or in the lungs, spine, or brain. Irregular clotting can be caused by heart problems, stroke, obesity, and smoking. Subtilisin enzymes can be used to break down potentially harmful clots. The protease nature of the enzyme can be used to hydrolyze the fibrin (an insoluble protein). Subtilisin blocks the binding site of the fibrin to prevent more clotting, as well as breaking down the peptide bonds that hold the protein together.
Design
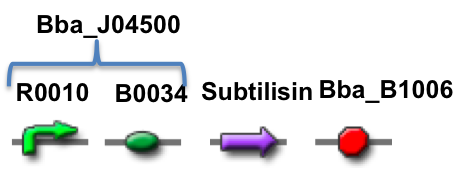
Our stain remover gene, subtilisin, is being synthesized with BioBrick prefix and suffix restriction site sequences. The gene has been delivered in a plasmid with the terminator sequence. It is depicted below.
(Promoter + RBS) + (BB-prefix + OmpA + Pro-sequence + Subtilisin + Terminator + BB-Suffix)
Basic Descriptions:
The promoter and RBS are together in BioBrick Part BBa_J04500, which is the same part used in the practice 3A assembly kit. BB-prefix is the BioBrick sequence with the appropriate restriction enzyme sites to conform to the 3A assembly standard. OmpA is a short signal peptide that coaxes E.coli to secrete the downstream protein out of the cell. Pro-sequence is a protein that helps guide the folding of the subtilisin enzyme and is part of the natural subtilisin gene. Subtilisin is a serine protease enzyme that degrade other proteins. BB-suffix is the BiobBick sequence with the appropriate restriction enzyme sites to conform to the 3A Assembly standard. The terminator is BBa_B1006.
Parts Explained:
The promoter is taken from the lac operon which is an inducible operon. A lac operon is a series of genes that code for proteins that digest lactose. However we have replaced those genes with genes that code for subtilisin. LacI codes for repressors that halt transcription because they prevent RNA polymerase from binding. It can be regulated because it is CAP sensitive. It has two binding sites, the first one allows the CAP protein to bind, while the second one binds to the LacI protein. The IPTG is a molecular mimic of lactose and the gene is turned on only when IPTG is present but when the glucose is absent, otherwise the gene cannot be turned on. RBS is the ribosome binding site, which is where the ribosomes bind together to begin translation. The goal is to translate a protein called subtilisin, which is a serine protein enzyme that is responsible for digesting protein molecules. Before the operon, there is a BB-prefix and after the termination sequence to cease transcription of subtilisin genes there is a BB-suffix. The two BB sequences are restriction sites where the the gene can be cut to be incorporated into a plasmid. The promoter and RBS together is called Part A (BBa_J04500) and that is 220 base pairs. So since the operon we are using is inducible, so when there is no glucose, there is an accumulation of cAMP which binds to CAP and which in turn allows transcription to occur. Also when when LacI is absent, the repressors are not produced and so transcription is possible. So if we want subtilisin to be produced then LacI should be absent, glucose should be absent but IPTG should be present.
I'll see of I can find pictures of the synthesized gene construct that was delivered from BioBasic.
OmpA signal peptide is 21 amino acids long. It helps to secrete recombinant enzymes out of the bacteria, where time is an important factor in how much has been secreted. Induction time requires several hours, 4 in the experiment. Normally, it is used to translocate and direct enzymes across the cell membrane of the bacteria; the type of bacteria should not be a major factor, but should also not be ignored. (The experiment used Gram-negative E. coli for over-expression of the test enzymes, using OmpA as the experimental.) For some enzymes, OmpA was equally effective in secretion as the native signal peptides, however this is not guaranteed for all enzymes. Due to the large variety of enzymes that OmpA can direct, it is advantageous to use it for non-native enzyme secretion in transformed bacteria.
I'll find a 3D crystal structure that highlights the active site of the enzyme.
Subtilisin is a protein-degrading enzyme. The peptide bonds are degraded with serine residue at the active site. The enzyme works in a signal-transduction pathway, with Ser-221 cleaving the peptide bond with its partially negative oxygen atom. In commercial use, subtilisin is genetically engineered because detergents and high temperatures easily inactivate the wild type. Subtilisin is used to remove protein based stains. It usually consists of 269 to 275 amino acids and its active from pH 6 to 11, with its major activity from pH 9 to 11. Subtilisin is easily found in the environment and is readily biodegradable. Subtilisin can cause a respiratory allergy in some people when come in contact with it in detergent. Reactions differ with the frequency, magnitude, and duration of the exposure. There is not a lower bench for the allergy currently. Because of the low concentrations in detergent, Subtilisin is not a concern for skin or eye irritation. There are no major health concerns for the use of Subtilisin in laundry and cleaning products. Subtilisin comes from the Bacillus genus. The overall tertiary structure of the species is the same, which means a similar function.
Procedure
Before we commenced the experiment, we worked through the practice round. Using the 3A Assembly and Transformation Efficiency Kit, we tested restriction digests, gel electrophoresis, ligation, and prokaryotic transformation.
Our experiment required the DNA of the subtilisin gene to be sequenced and synthesized. Using the company BioBasics, 10 ul of the resulting DNA was created for the transformation.
Results
Human Practices
Safety
E-Notebook
Sponsors
The Waltham Education and Beyond Foundation and the Sally Elizabeth Peters Enrichment Program provided a grant to help us fund the lab work and iGEM Jamboree.
New England BioLabs generously donated enzymes and competent cells to support our project.
Bio Basics Inc. supplied us with synthesized DNA.
 "
"