Team:Montgomery Cougars NJUSA/Project
From 2014hs.igem.org
(→Project Overview) |
(→Results and Conclusion) |
||
| Line 49: | Line 49: | ||
==Results and Conclusion== | ==Results and Conclusion== | ||
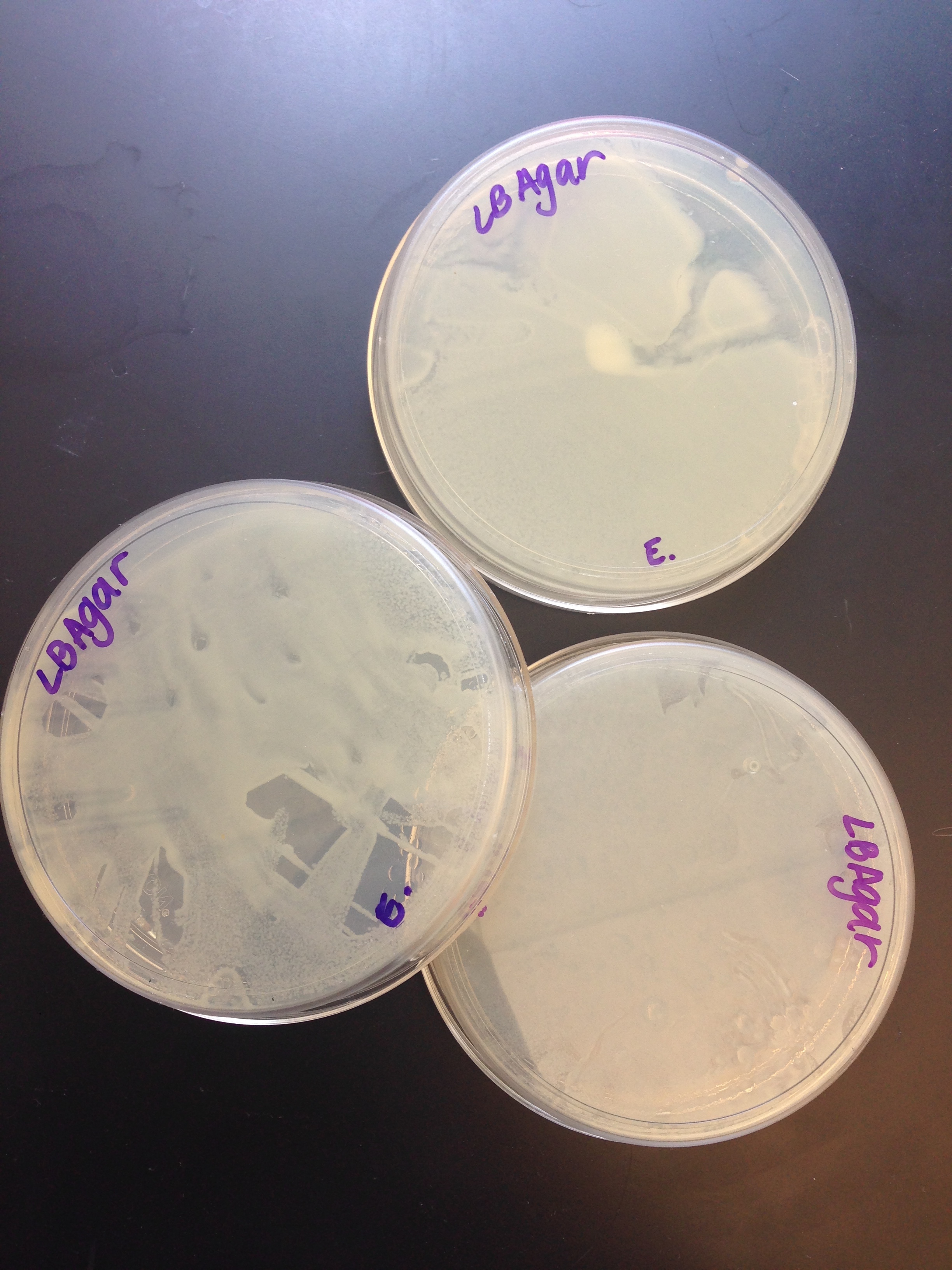
| - | [[Image:Growth.jpg| | + | [[Image:Growth.jpg|right|300px|thumb|Our completely grown transformed bacteria)]] |
While we were extremely hopeful that our lab would work, our bacteria unfortunately did not glow. This indicates that our mechanism was ultimately unsuccessful. When we checked on our bacteria cultures, we found that the bacteria did indeed grow but did not display the correct reaction when viewed under a UV light. We can only conclude that our bacteria either did not accept the plasmid, the plasmid was ligated incorrectly or the promotor was not stimulated. Our suspicions lie in the fact that our promotor was not stimulated. We realized that the lacl regulated promotor would not be initiated in the LB Agar plate. However, due to our limited time, we could not obtain what was needed for our project. Also, during our transformation process, we had to forego the RFP control in order to finish the experiment in time. Our ligation procedure was much more complex than the procedure given, so we had to experiment and create our own ligation process. All in all, our first year laboratory was very shaky and our lack of experience forced us to take many risks. The lack of professional laboratory setting could also have contributed to our failure to create our desired protein. | While we were extremely hopeful that our lab would work, our bacteria unfortunately did not glow. This indicates that our mechanism was ultimately unsuccessful. When we checked on our bacteria cultures, we found that the bacteria did indeed grow but did not display the correct reaction when viewed under a UV light. We can only conclude that our bacteria either did not accept the plasmid, the plasmid was ligated incorrectly or the promotor was not stimulated. Our suspicions lie in the fact that our promotor was not stimulated. We realized that the lacl regulated promotor would not be initiated in the LB Agar plate. However, due to our limited time, we could not obtain what was needed for our project. Also, during our transformation process, we had to forego the RFP control in order to finish the experiment in time. Our ligation procedure was much more complex than the procedure given, so we had to experiment and create our own ligation process. All in all, our first year laboratory was very shaky and our lack of experience forced us to take many risks. The lack of professional laboratory setting could also have contributed to our failure to create our desired protein. | ||
Revision as of 20:12, 20 June 2014
Contents |
Project Overview
The Role of P. Acnes in Acne Vulgaris
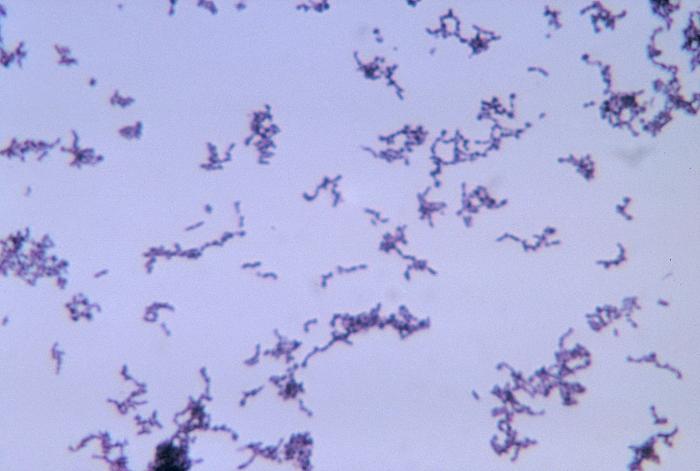
The bacteria Propionibacterium acnes is responsible for Acne Vulgaris, more commonly known as acne. Although P. Acnes is not an issue for most people, there are certain virulent strains that exacerbate acne inflammation. The bacteria thrives in the presence of lipids, specifically sebum, which serves as the primary source of the bacteria's energy and sustenance. An increase of sebum production attracts bacteria into hair follicles, where the bacteria can multiply and colonize, resulting in an inflammatory reaction of cysts and pustules, which can lead to acne scars if not treated properly.
Description
For the 2014 Competition season, Montgomery_Cougars_NJUSA aims to create a better solution for acne reduction. Acne is a problem that affects nearly 80% of American adolescents. Unfortunately, this widespread problem does not have a concrete solution. Leading acne medication companies, such as Proactiv, use benzyl peroxide and/or salicylic acid alike in their products, in which the harsh chemicals often shed the skin. Although this method may unclog pores from the patient’s face, it is moderately painful and is not always 100% effective. Therefore, to combat this widespread ailment, Montgomery iGEM has decided to take a different approach.
In an effort to create a less invasive treatment, we decided to design an enzyme to break down the sebum on human skin, thereby reducing the bacteria's source of energy and ability to colonize. Because sebum is made of triglyceride oils, wax esters, squalene, and metabolites of fat-producing cells, we sought a group of enzymes that would break down these components. Through our research, we have observed a definite positive correlation between patients with acne and presence of triglycerides and wax esters. Our mechanism ultimately focused on targeting these substances. In reducing these substances, we effectively deplete the food that the P. acne bacteria needs to thrive and elicit the inflammatory response we perceive to be acne.
Enzyme/Gene Selection: The Process
When we first started our product research, we had many complications to address. At first, we planned on creating an antibiotic that would directly combat the bacteria. Once we realized that designing a bacteria, namely E. Coli, to synthesize a protein to break down other bacteria was much beyond our scope, we decided to focus on what exactly makes the bacteria thrive. Thus, we moved onto examining the sebum within the skin that allows bacteria to grow and indirectly create the inflammatory response known as acne. As mentioned before, sebum is composed of wax esters, triglycerides, squalene among other lipids and fats native to the skin of humans. We hypothesized that if we limited the food supply for the bacteria, the persistence of acne would decrease.
Finding a way to break down the components of squalene was a challenge in itself. Our research revealed that sebum is primarily broken down through beta oxidation, which involves the removal of hydrogen by dehydrogenase enzymes. However, it was extremely difficult to find a suitable dehydrogenase for our mechanism. The most prevalent component of sebum is sapienic acid, which plays a key role in the formation of acne. Sapienic acid is difficult to break down because of its carboxylic functional group, which is rarely targeted by the body's natural array of enzymes. We unsuccessfully tried to find an enzyme to target carboxyls. Our breakthrough came in late May, when our research brought us to the conclusion that another component of sebum, wax esters, could be broken down by the method we had previously proposed: alcohol dehydrogenase. Wax esters contain alcohol functional groups that our mechanism targets. Thus, at least theoretically, we have created an alternative method to reduce acne production in humans.
Final Design
Our team decided to create two biological systems, each designed to produce a specific protein. The two engineered plasmids contain the same promoter, RBS, and terminator; the only part that differs is the actual Protein Coding Region. One plasmid will produce the Medium Chain Alcohol Dehydrogenase; the other will produce the Medium Chain Aldehyde Dehydrogenase. The function of both of these proteins is to change the shape of the wax esters, a central component of sebum, and thus limit the P. acnes' nutrient source.
Wax esters contain both alcohol and aldehyde groups, which are the targets of the enzymes that we plan to express.
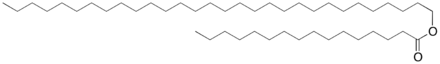
Our main purpose for both of these plasmids is to harvest a protein, so we needed to choose parts that would ensure a maximum efficiency rate.
The promoter chosen is the naturally found LacZYA operon in E.coli (BBa_R0010). Because it is an inducible, multiregulated operon (positive CAP control and Lacl negative control), using this promoter can give us a higher level of regulation over the plasmid. We may be able to control transcription efficiency rates by changing the environment that the E.coli colonies grow in. The Elowitz RBS (BBa_B0034) is the definition of efficiency 1.0, as tested by team Warsaw in 2010. We chose this RBS to maximize the translation efficiency rate that will produce the protein of interest. We chose a double terminator (BBa_B0015) that would ensure high efficiency in terminating transcription, and less error when we harvest the protein.
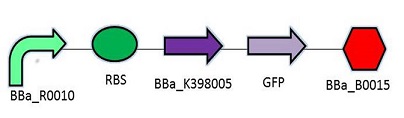
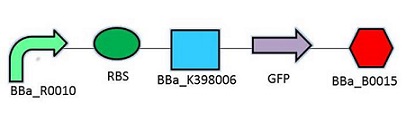
Promotor: BBa_R0010
RBS: BBa_B0034
Protein Coding Domain: BBa_K398005 or BBa_K398006
Signal Protein: BBa_E0040
Terminator: BBa_B0015
Results and Conclusion
While we were extremely hopeful that our lab would work, our bacteria unfortunately did not glow. This indicates that our mechanism was ultimately unsuccessful. When we checked on our bacteria cultures, we found that the bacteria did indeed grow but did not display the correct reaction when viewed under a UV light. We can only conclude that our bacteria either did not accept the plasmid, the plasmid was ligated incorrectly or the promotor was not stimulated. Our suspicions lie in the fact that our promotor was not stimulated. We realized that the lacl regulated promotor would not be initiated in the LB Agar plate. However, due to our limited time, we could not obtain what was needed for our project. Also, during our transformation process, we had to forego the RFP control in order to finish the experiment in time. Our ligation procedure was much more complex than the procedure given, so we had to experiment and create our own ligation process. All in all, our first year laboratory was very shaky and our lack of experience forced us to take many risks. The lack of professional laboratory setting could also have contributed to our failure to create our desired protein.
We hold in our belief that our project, when done correctly could have remedied the acne that plagues thousands of adolescents in America. We hope that next year, our endeavors are more successful.
 "
"