Team:CIDEB-UANL Mexico/project capture
From 2014hs.igem.org
| Line 47: | Line 47: | ||
<p>After the removal of sodium ions from salty water, it is needed to remove Cl- ions as well in order to complete the desalination process. E. CARU only captures Na+ ions, leaving Cl- ions in the water medium; but since Cl is a diatomic molecule (meaning it cannot be alone in normal conditions), it joins another Cl molecule, forming Cl2.</p> | <p>After the removal of sodium ions from salty water, it is needed to remove Cl- ions as well in order to complete the desalination process. E. CARU only captures Na+ ions, leaving Cl- ions in the water medium; but since Cl is a diatomic molecule (meaning it cannot be alone in normal conditions), it joins another Cl molecule, forming Cl2.</p> | ||
<p>Normally, Cl2 is in a gaseous state at normal conditions, so what would remain after E. CARU takes Na+ ions from water is a mixture of Cl2 gas and water molecules. In order to remove it from water it is possible to use a method involving the separation of a gas from a liquid based in the boiling points of each component in the mixture. Cl2 gas has a boiling point of -34.6°C and water has a boiling point of 100°C. By cooling the mixture at -34.6°C, the gas would evaporate, separating itself from water. (Bentor, 2014).</p> | <p>Normally, Cl2 is in a gaseous state at normal conditions, so what would remain after E. CARU takes Na+ ions from water is a mixture of Cl2 gas and water molecules. In order to remove it from water it is possible to use a method involving the separation of a gas from a liquid based in the boiling points of each component in the mixture. Cl2 gas has a boiling point of -34.6°C and water has a boiling point of 100°C. By cooling the mixture at -34.6°C, the gas would evaporate, separating itself from water. (Bentor, 2014).</p> | ||
| - | <p>Yet still, there is an important factor to consider: Cl2 gas is toxic, but as it is 2.5 times heavier than air (CFC StarTec LLC, 2007), it would stay in water at room temperature. For this reason, before cooling the Cl2 gas in order to take it away, it is necessary to take safety measurements. The one proposed by the team is the usage of an Atomic Absorption Spectroscopy (see | + | <p>Yet still, there is an important factor to consider: Cl2 gas is toxic, but as it is 2.5 times heavier than air (CFC StarTec LLC, 2007), it would stay in water at room temperature. For this reason, before cooling the Cl2 gas in order to take it away, it is necessary to take safety measurements. The one proposed by the team is the usage of an Atomic Absorption Spectroscopy (see <b>Figure 2</b>). </p> |
<p>Atomic Absorption Spectrometry (AAS) is an analytical technique that measures the concentrations of elements. Atomic absorption is so sensitive that it can measure down to parts per billion of a gram (µg dm–3). The technique uses light wavelengths specifically absorbed by an element that correspond to the energies needed to excite electrons from one energy level to a higher one. (Royal Society of Chemistry).</p> | <p>Atomic Absorption Spectrometry (AAS) is an analytical technique that measures the concentrations of elements. Atomic absorption is so sensitive that it can measure down to parts per billion of a gram (µg dm–3). The technique uses light wavelengths specifically absorbed by an element that correspond to the energies needed to excite electrons from one energy level to a higher one. (Royal Society of Chemistry).</p> | ||
| Line 62: | Line 62: | ||
<p>It is important to be familiarized with what it is being worked with, and since this putative protein has never been used at iGEM before, it was done a lot of research on it. </p> | <p>It is important to be familiarized with what it is being worked with, and since this putative protein has never been used at iGEM before, it was done a lot of research on it. </p> | ||
| - | <p>The composition and form of a protein show relevant data about its actions and functions, that it is why it was investigated NhaS’ predicted type. It was found in the modelling tool <a href="http://bmm.cancerresearchuk.org/~3djigsaw/" target="_blank">3D-JIGSAW</a> from <i>Cancer Research UK'</i> site that its possible protein or peptide type would be helix, coil or strand <b>(Figure | + | <p>The composition and form of a protein show relevant data about its actions and functions, that it is why it was investigated NhaS’ predicted type. It was found in the modelling tool <a href="http://bmm.cancerresearchuk.org/~3djigsaw/" target="_blank">3D-JIGSAW</a> from <i>Cancer Research UK'</i> site that its possible protein or peptide type would be helix, coil or strand <b>(Figure 3)</b>. Also, it was found in the <a href="http://raptorx.uchicago.edu/StructurePrediction/myjobs/30991450_67869/" target="_blank"> <i>Raptor X</i></a> protein modelling site a predicted structure of the protein. <b>(Figure 4)</b></p> |
<center> | <center> | ||
| Line 68: | Line 68: | ||
<p><img width=677 height=50 src="https://static.igem.org/mediawiki/2014hs/5/54/PPRESULTS.jpg"align=center hspace=12 alt="IMG_0317"></p> | <p><img width=677 height=50 src="https://static.igem.org/mediawiki/2014hs/5/54/PPRESULTS.jpg"align=center hspace=12 alt="IMG_0317"></p> | ||
| - | <p><b>Figure | + | <p><b>Figure 3.</b> Interactive 3D-JIGSAW's result that indicates the predicted protein type of NhaS.</p> |
<br><br> | <br><br> | ||
<p><img width=780 height=170 src="https://static.igem.org/mediawiki/2014hs/7/7c/NhaS_proteinCIDEB.jpg"align=center hspace=12 alt="IMG_0317"></p> | <p><img width=780 height=170 src="https://static.igem.org/mediawiki/2014hs/7/7c/NhaS_proteinCIDEB.jpg"align=center hspace=12 alt="IMG_0317"></p> | ||
| - | <p><b>Figure | + | <p><b>Figure 4.</b> Predicted protein structural results from <i>Raptor X</i>.</p></center><br> |
| - | <p>Research was done in different sources. The previous information was confirmed in <i>Predict Protein</i> site. As shown in <b>Figure | + | <p>Research was done in different sources. The previous information was confirmed in <i>Predict Protein</i> site. As shown in <b>Figure 5</b> and <b>Figure 6</b>.</p><br><center> |
<p><img width=561 height=239 src="https://static.igem.org/mediawiki/2014hs/9/98/GraphsCIDEB.jpg"align=center hspace=12 alt="IMG_0317"></p> | <p><img width=561 height=239 src="https://static.igem.org/mediawiki/2014hs/9/98/GraphsCIDEB.jpg"align=center hspace=12 alt="IMG_0317"></p> | ||
| - | <p><b>Figure | + | <p><b>Figure 5.</b> Results given by <i>Predict Protein</i> showing the </br>secondary structure composition and solvent accessibility of the putative NhaS protein.</p></center><center> |
<br><br> | <br><br> | ||
<p><img width=796 height=171 src="https://static.igem.org/mediawiki/2014hs/5/5b/PPCIDEB.jpg"align=center hspace=12 alt="IMG_0317"></p> | <p><img width=796 height=171 src="https://static.igem.org/mediawiki/2014hs/5/5b/PPCIDEB.jpg"align=center hspace=12 alt="IMG_0317"></p> | ||
| - | <p><b>Figure | + | <p><b>Figure 6.</b> Results given by <i>Predict Protein</i> showing the predicted precise structure of the NhaS protein. </p></center> |
<br> | <br> | ||
<p>As it can be apprecciated, the results from all sources match, so, based on the previous information it was concluded that NhaS is most possible to be of the helix type. Being aware of the secondary structure of proteins is relevant, since hence, the protein folding mechanism can be taken into consideration.</p> | <p>As it can be apprecciated, the results from all sources match, so, based on the previous information it was concluded that NhaS is most possible to be of the helix type. Being aware of the secondary structure of proteins is relevant, since hence, the protein folding mechanism can be taken into consideration.</p> | ||
| - | <p>According to Krulwich and Ivey (1994), the location of the protein is in the cytoplasmic side of the membrane, however, when the research was made, it was found out that the protein is predicted to be highly non-cytoplasmic (Stockholm Bioinformatics Center SBC Phobius, 2014), as shown in <b>Figure | + | <p>According to Krulwich and Ivey (1994), the location of the protein is in the cytoplasmic side of the membrane, however, when the research was made, it was found out that the protein is predicted to be highly non-cytoplasmic (Stockholm Bioinformatics Center SBC Phobius, 2014), as shown in <b>Figure 7</b>. Based on this, the team came out with a hypothesis in which NhaS would be located in the inner part of the membrane but on its cytoplasmic side, and since it is a helix protein, some parts would be exposed and some would not. This would explain both predictions of both sources of information.</p><center><br> |
<p><img width=550 height=420 src="https://static.igem.org/mediawiki/2014hs/3/32/ProteinCIDEB.jpg" align=center hspace=12 alt="IMG_0317"></p> | <p><img width=550 height=420 src="https://static.igem.org/mediawiki/2014hs/3/32/ProteinCIDEB.jpg" align=center hspace=12 alt="IMG_0317"></p> | ||
| - | <p><b>Figure | + | <p><b>Figure 7.</b> Predicted protein overview results from <i>Stockholm Bioinformatics Center SBC Phobius</i>.</p> |
</center> | </center> | ||
| Line 100: | Line 100: | ||
<p><img width=250 height=200 src="https://static.igem.org/mediawiki/2014hs/2/28/PPCIDEB2.jpg"align=center hspace=12 alt="IMG_0317"></p> | <p><img width=250 height=200 src="https://static.igem.org/mediawiki/2014hs/2/28/PPCIDEB2.jpg"align=center hspace=12 alt="IMG_0317"></p> | ||
| - | <p><b>Figure | + | <p><b>Figure 8.</b>“Predicted localization for the Bacteria domain: Inner Membrane (GO term ID: 0005886) Prediction confidence 76.”</p></center> |
<br> | <br> | ||
| Line 123: | Line 123: | ||
<center> | <center> | ||
<p><img width=460 height=250 src="https://static.igem.org/mediawiki/2014hs/3/32/TestCaptureAndAroma.jpg" align=center hspace=12></p> | <p><img width=460 height=250 src="https://static.igem.org/mediawiki/2014hs/3/32/TestCaptureAndAroma.jpg" align=center hspace=12></p> | ||
| - | <p><b>Figure | + | <p><b>Figure 9.</b> Project's ideal device and how it was divided for its respective tests.</p></center> |
<br> | <br> | ||
Revision as of 18:20, 20 June 2014
Capture Module

As a step to desalinize water, iGEM CIDEB 2014's project intended to capture sodium ions from saline water using a protein named NhaS produced from the nhaS gene expression. This gene was obtained from a 1994 patent by Krulwich & Ivey and no further information was found about the gene in other sources. A lot of research of related information and predictions about the NhaS protein in different sites had to be made in order to obtain more information about it and being able to work with and include it in the project.
Description
NhaS is a putative protein from Bacillus firmus that is characterized by its ability to bind and sequestering sodium ions, with a calculated weight of 7100 Daltons and a pH of 12. It “can enhance the Na+ -resistance of antiporter- deficient strains by increasing the availability of Na+ to the integral membrane antiporters on the cytoplasmic side of the membrane and by sequestering Na+ while rate-limiting efflux mechanisms catalyze extrusion of the cation.” (Krulwich & Ivey, 1994)
Research by Krulwich and Ivey (1994) supports that in its origin bacteria, NhaS works as a regulation pH homeostasis protein because it makes the cytoplasmic pH more acidic than the external medium, usually basic.
Essentially, NhaS performs three different functions; (1) capturing sodium ions, (2) regulating pH, and (3) enhancing the resistance of bacteria to high saline conditions.

Figure 1. Patent US 5346815 A shows extracts of the E. coli EP432 transformed with pGEM (fig. 4A) and pGRVH (fig. 4B). pGEM is a control plasmid and pGRVH is a plasmid with the nhaS gene. Those are crude extracts shown by the effect of putting the bacteria to an SFBI excitation, which is a sodium-sensitive molecule used to measure intracellular Na+. Resuming, it shows in basic draws that the protein is expressed in E. coli and in what quantity according to the excitation level where it is exposed.
What would happen with Cl- ions?
After the removal of sodium ions from salty water, it is needed to remove Cl- ions as well in order to complete the desalination process. E. CARU only captures Na+ ions, leaving Cl- ions in the water medium; but since Cl is a diatomic molecule (meaning it cannot be alone in normal conditions), it joins another Cl molecule, forming Cl2.
Normally, Cl2 is in a gaseous state at normal conditions, so what would remain after E. CARU takes Na+ ions from water is a mixture of Cl2 gas and water molecules. In order to remove it from water it is possible to use a method involving the separation of a gas from a liquid based in the boiling points of each component in the mixture. Cl2 gas has a boiling point of -34.6°C and water has a boiling point of 100°C. By cooling the mixture at -34.6°C, the gas would evaporate, separating itself from water. (Bentor, 2014).
Yet still, there is an important factor to consider: Cl2 gas is toxic, but as it is 2.5 times heavier than air (CFC StarTec LLC, 2007), it would stay in water at room temperature. For this reason, before cooling the Cl2 gas in order to take it away, it is necessary to take safety measurements. The one proposed by the team is the usage of an Atomic Absorption Spectroscopy (see Figure 2).
Atomic Absorption Spectrometry (AAS) is an analytical technique that measures the concentrations of elements. Atomic absorption is so sensitive that it can measure down to parts per billion of a gram (µg dm–3). The technique uses light wavelengths specifically absorbed by an element that correspond to the energies needed to excite electrons from one energy level to a higher one. (Royal Society of Chemistry).

Figure 2. Atomic Absorption Spectroscopy dispositive.
The use of an Atomic Absorption Spectrometry is a way to remove the Cl2 gas from water and measure it to prevent its escape. Then, the stored Cl2 gas can be used to sterilize drinking water, to disinfect swimming pools and to be part in the manufacture of many consumer products; such as paper, dyestuffs, textiles, petroleum products, medicines, antiseptics, insecticides, foodstuffs, solvents, paints, and plastics (especially PVC). It can also be used to produce bleaches, chlorates, chloroform, carbon tetrachloride and bromine. A further substantial use for this element is in organic chemistry, both as an oxidizing agent and in substitution reactions (Emsley, 2011).
Research on NhaS
It is important to be familiarized with what it is being worked with, and since this putative protein has never been used at iGEM before, it was done a lot of research on it.
The composition and form of a protein show relevant data about its actions and functions, that it is why it was investigated NhaS’ predicted type. It was found in the modelling tool 3D-JIGSAW from Cancer Research UK' site that its possible protein or peptide type would be helix, coil or strand (Figure 3). Also, it was found in the Raptor X protein modelling site a predicted structure of the protein. (Figure 4)

Figure 3. Interactive 3D-JIGSAW's result that indicates the predicted protein type of NhaS.

Figure 4. Predicted protein structural results from Raptor X.
Research was done in different sources. The previous information was confirmed in Predict Protein site. As shown in Figure 5 and Figure 6.

Figure 5. Results given by Predict Protein showing the secondary structure composition and solvent accessibility of the putative NhaS protein.

Figure 6. Results given by Predict Protein showing the predicted precise structure of the NhaS protein.
As it can be apprecciated, the results from all sources match, so, based on the previous information it was concluded that NhaS is most possible to be of the helix type. Being aware of the secondary structure of proteins is relevant, since hence, the protein folding mechanism can be taken into consideration.
According to Krulwich and Ivey (1994), the location of the protein is in the cytoplasmic side of the membrane, however, when the research was made, it was found out that the protein is predicted to be highly non-cytoplasmic (Stockholm Bioinformatics Center SBC Phobius, 2014), as shown in Figure 7. Based on this, the team came out with a hypothesis in which NhaS would be located in the inner part of the membrane but on its cytoplasmic side, and since it is a helix protein, some parts would be exposed and some would not. This would explain both predictions of both sources of information.

Figure 7. Predicted protein overview results from Stockholm Bioinformatics Center SBC Phobius.
As a “confirmation” for the previous hypothesis made, Predict Protein site gave the following result:
Figure 8.“Predicted localization for the Bacteria domain: Inner Membrane (GO term ID: 0005886) Prediction confidence 76.”
Based on the predicted result and with the previous hypothesis the team formulated, it was concluded that the protein would act in the cytoplasmic side of the inner membrane. This information was used for the understanding and explaining of the module, as well as for designing different animations.
Further information about NhaS and the experimental results made by the team about it can be found in its parts registry section, as well as the results section in this wiki.
Device
The proposed circuit for the module is as follows: An UV promoter, an RBS; the nhaS gene, another RBS, a WinterGreen odor reporter (BSMT1 Opt.) and a terminator.
However, for practical and experimental reasons explained in the next paragraph, it was decided that nhaS and BSMT1 Opt. would be tested separately, leading to both Capture and Aroma modules.
As mentioned before, in the ideal project it is proposed an odor Wintergreen (BSMT1 Opt.) reporter for this module in order to know if there is any production and expression of NhaS, but since BSMT1 Opt. wasn't proved yet and NhaS is a putative protein it was decided to test NhaS physically with RFP, which is a simple and common reporter, in order to observe functional results in the tests for this specific segment of the project, obtaining conclusions about both genes, and being able to put them together in the ideal device.
Further information about the odor reporter can be found in the Aroma module in this wiki.
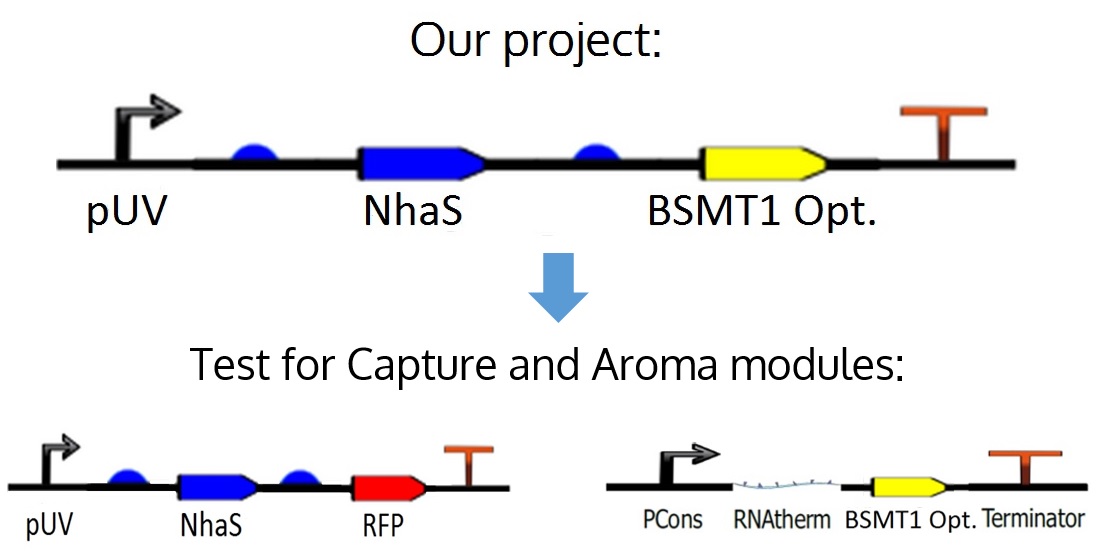
Figure 9. Project's ideal device and how it was divided for its respective tests.
Capture module's parts description
|
IMAGE |
CODE |
DESCRIPTION |
|
|
UV Promoter from iGEM Colombian Team 2007. Its length is 76bp.
Tim ITB_Indonesia 2013 proved that this part worked for them. |
|
|
|
Very common Ribosome Binding Site, based on Elowitz repressilator. Its length is 12bp
.
|
|
|
|
|
nhaS is a putative gene that produces the NhaS protein. iGEM CIDEB 2014 introduced it to iGEM for the first time. Its length is 207bp. |
|
|
|
Wintergreen-odor enzyme generator, used to allow the production of methyl salicylate, when it is induced by salicilyc acid. Its length is 1074bp. |
|
|
Part made of 6pb responsible for stopping transcription.
|
Justifications
UV promoter as an initiator for the system
|
|
The NhaS’ production and functionability was regulated using a promoter from iGEM Colombian Team 2007 that is regulated by UV irradiation and does not causes mutations in the bacteria. This type of promoter was chosen in order to have a control about the NhaS action in the project, so it would only be activated under UV rays, and therefore, it could be decided at which time the NhaS gene was going to be expressed. There were chosen UV rays as the initiator of the expression because they are present in normal conditions, and in case the biofilter was used in a water treatment plant, as it is intended as a future projection for the project, it could be simpler and cheaper to activate the system with sunlight, since the mentioned promoter is activated under 360 wavelengths of light spectrum. As an extra, the activation of the system without the use of reactants that could pollute the water is one of the advantages of its use. |
Other teams that used pUV and NhaS
In the case of pUV
Before choosing the BBa_I765001 UV promoter the team reviewed the experiences by other participants with it, with the intention of assuring the maximum possible success rate if used in this module. The following table shows information about its usage with other teams.
|
Team |
Use |
|
An UV induced promoter and a CDS of a testosterone-making gene.
|
|
|
|
pUV and mCherry (BBa_J06702) to detect DNA damage.
|
|
|
Expression of the UV promoter in presence of UV irradiation light lead to the expression of the EYFP reporter. |
In the case of nhaS
There are no teams that used this gene in the past. IGEM CIDEB 2014 was the first team that synthesized, characterized, tested and registered it in the parts registry.
Future usage
The team considers NhaS to be useful for the scientific community in general, in addition to iGEM. NhaS, as mentioned before, is a putative protein. Knowing specific results about its functions could be of great use for future projects.
The potential for future usage of this gene is great. As mentioned in the patent from Krulwich and Ivey (1994), the gene encoding NhaS can be introduced into cells to produce desalination bioreactors, can be introduced into plants as a transgene to produce plants that are resistant to sodium, may be used for treatments involving NNa+ /K+ ATPase disorders, e.g., in heart disease, and may be introduced parenterally, preferably orally, to bind to and sequester dietary sodium. The possibilities are wide and promising.
Capture Module Zoom In
Bibliography/References
● Antiquity. (2013, January 31). Part: BBa_B0034. Retrieved May 1st, 2014, from http://parts.igem.org/wiki/index.php?title=Part:BBa_B0034
● Bentor, Y. (18 de Jun de 2014). Chlorine. Obtained from Chemicalelement: http://www.chemicalelements.com/elements/cl.html
● BMM Cancer Research UK. Interactive 3D Jigsaw. Retrieved May 1st, 2014, from bmm.cancerresearchuk.org
● CFC StarTec LLC. (24 de Dec de 2007). Chlorine Cl2. Obtained from CFC StarTec LLC: http://www.c-f-c.com/specgas_products/chlorine.htm
● Colombia Israel Team. (2007). Part: BBa_I765001. Retrieved April 1st, 2014, from Part-BBa_I765001
● Emsley, J. (2011). Chlorine. Obtained from Periodic table: http://www.rsc.org/periodic-table/element/17/www.matvalue.com
● IGEM Colombian Team. (2007). UV promoter. Retrieved April 1st, 2014, from http://parts.igem.org/Part:BBa_I765001
● Jianzhu Ma, Sheng Wang, Feng Zhao, and Jinbo Xu. Protein threading using context-specific alignment potential. Bioinformatics (Proceedings of ISMB 2013), Vol. 29, Issue 13, pp. i257-i265. From http://bioinformatics.oxfordjournals.org/content/29/13/i257.full/
● Jian Peng and Jinbo Xu. A multiple-template approach to protein threading. PROTEINS, 2011 Jun;79(6):1930-9. doi: 10.1002/prot.23016. Epub 2011 Apr 4. From http://www.ncbi.nlm.nih.gov/pubmed/21465564/
● Jian Peng and Jinbo Xu. RaptorX: exploiting structure information for protein alignment by statistical inference. PROTEINS, 2011, Vol 79, Issue S10, pp. 161-171. From
● Knight Lab. (2006). Part: BBa_B1002. Retrieved from http://parts.igem.org/Part:BBa_B1002 ● KRULWICH Terry, IVEY Mark.(1994) Sodium ion binding proteins. Retrieved April 1st, 2014, from http://www.google.com.mx/patents/US5346815 ● Morten Källberg, Haipeng Wang, Sheng Wang, Jian Peng, Zhiyong Wang, Hui Lu, and Jinbo Xu. Template-based protein structure modeling using the RaptorX web server. Nature Protocols 7, 1511-1522, 2012. From http://www.nature.com/nprot/journal/v7/n8/full/nprot.2012.085.html/ ● Predict Protein. Request ID: 473277. Retrieved May 3rd, 2014, from https://www.predictprotein.org ● Raptor X (2014). Protein Structure and Function Prediction: Result for job NhaS. Retrieved June 18th, 2014, from http://raptorx.uchicago.edu/StructurePrediction/myjobs/30991450_67869/ ● Royal Society of Chemistry. (s.f.). Atomic absorption spechtrometry. London: Burlington House. ● Stockholm Bioinformatics Center SBC (2014). Phobius. Retrieved from http://phobius.sbc.su.se
 "
"


















